The world is quickly advancing toward technology. Most individuals and organizations have adopted automation solutions, while others have embraced the idea of having smart home systems. These technological solutions require exports to develop, troubleshoot, monitor, and maintain. For this reason, careers in the tech space have grown tremendously, with many students opting for computer science-affiliated courses. The latter learning programs are more marketable, and the job market is always ready to absorb more skilled graduates. However, the popularity of computer science and information technology courses has led to a rise in the number and complexity of coding assignments to ensure students get skills that match the competitive job market.
Excelling in the computer science and information technology industry requires extra dedication to work. As a student, you must avail yourself to conduct extensive research to be able to learn and apply programming languages efficiently. Looking at the task, it takes work to keep up with everything. Therefore, you must be ready to experience fatigue due to endless sleepless nights and long sitting hours as you work on college assignments and side projects. To be a competent computer programmer, you must learn, understand, and master different computer programming languages, such as the unified modeling language (UML), HTML, Java, Python, and Go. While many students strive to balance life and academics, it is an uphill task for some people, mainly when there are other responsibilities. At this point, the only solution seems to find assistance from the best programming homework help websites to ensure you submit the project on time.
While students have been able to find coding homework help online, there is a challenge when it comes to choosing a trustworthy and reliable site. There are numerous factors to consider before you pay someone to do your programming assignment. In this article, we have narrowed down the best websites to go to when looking for legitimate online programming assignment help.
List of 5 Best Programming Assignment Help Websites
Suppose you have challenges mastering the core programming concepts or completing the assignments on time. In that case, you can consider programming homework help sites to offer you solutions at a relatively affordable price. Here is a list of top academic writing websites that can be of assistance:
- CodingHomeworkHelp — professional assistance with programming tasks
- DoMyAssignments — reliable STEM homework assistance
- AssignCode — excellent “write code for me” service
- CWAssignments — affordable and reliable “do my programming assignment” service
- CodingPedia — high speed delivery for your coding assignments
Detailed overview of the top programming homework help websites for college students
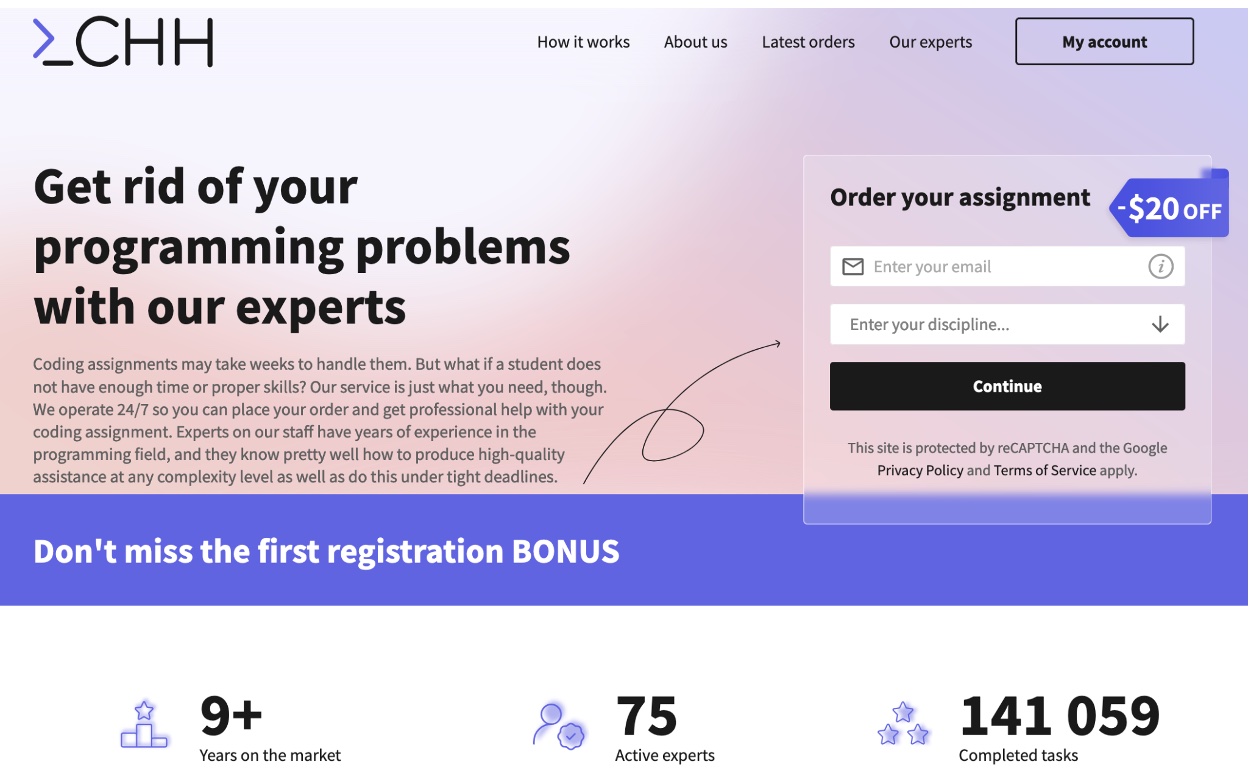
Codinghomeworkhelp.org – A top-notch programming assignment help website for college students
The company provides a wide range of services and features that make it stand out as one of the best resources for struggling students with their programming assignments.
One of the most significant features of CodingHomeworkHelp is the quality of its tutors. The website boasts a team of highly qualified and experienced tutors who are experts in various programming languages and disciplines. These tutors deeply understand the material and can provide students with detailed and accurate answers to their questions. They are also very responsive, providing quick and helpful answers to any questions or concerns that students may have. This attribute benefits students who need help understanding complex concepts or troubleshooting code.
In addition to its great tutors, codinghomeworkhelp.org also offers a user-friendly interface that makes it easy for students to find and access the help they need. The website is well-organized and easy to navigate, with a wide range of resources and tutorials available to help students learn and understand the material. Also, codinghomeworkhelp.org tutors are available around the clock. The 24-hour availability means that students can get help whenever they need it, regardless of their time zone or schedule. The arrangement is beneficial for students who have busy schedules or are working on assignments outside of class and allows them to get the help they need, even if they have to work late into the night or on weekends. The flexible schedule is a huge advantage for students needing help to keep up with the demands of their coursework.
Lastly, codinghomeworkhelp.org provides plagiarism-free work and a timely delivery guarantee. These are essential aspects that students should consider since all college papers have a fixed timeline. In addition, the company provides a money-back guarantee in case a student is unsatisfied with their service. The features present it as an ideal company for seeking coding homework help. Therefore, this is a good one if you are stranded for a choice.
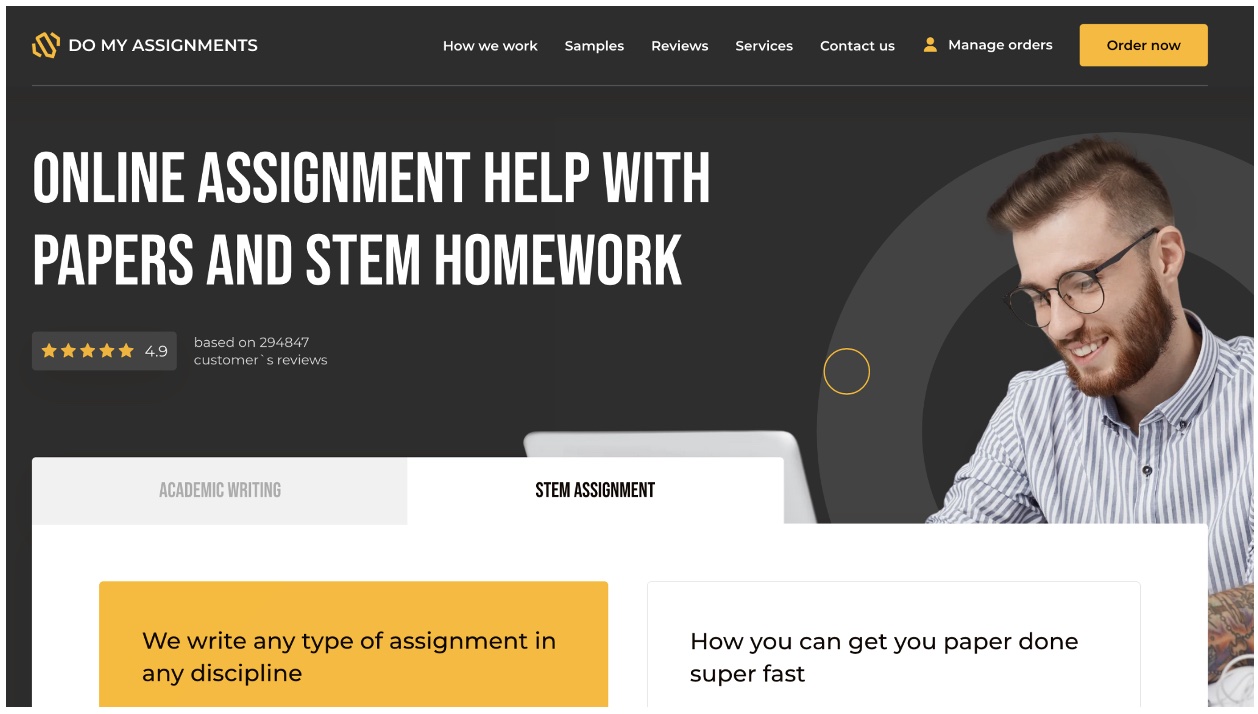
DoMyAssignments.com – An outstanding coding assignment help service for college students
The company is dedicated to providing the standard features that make it a darling to learners struggling with complex programming assignments. The company has programmers who deeply understand the material and can provide students with detailed and accurate answers to their questions. They are also very patient and approachable, explaining and working through problems with students until they fully understand the concepts.
DoMyAssignments has one of the most user-friendly interfaces. The feature makes it easy for students to find and access the help they need. In addition, it is well-organized and intuitively designed, with a wide range of resources and tutorials available to help students learn and understand the material. The features and resources include a comprehensive FAQ section and a library of sample assignments which help to build the student’s confidence in the company’s ability.
In addition, the company has set itself apart from other programming assignment help websites regarding their quality of work, timely delivery, and plagiarism-free work. The quality assurance team implements strict policies to ensure the expert’s hand in the highest quality work will help the students get the best grades. The site also has a money-back guarantee, a limited revision policy, and the privacy policy to ensure the clients get the service they deserve.
The last significant aspect of the company is the support team. DoMyAssignments.com’s customer service team is available 24/7, so students can get help whenever they need it, regardless of their time zone or schedule. The schedule is helpful to students from different time zones and those that have to sit for extra long hours trying to understand the complex programming concepts typical of the college curriculum. As a result, the company stands out as an ideal helper to students seeking coding help.
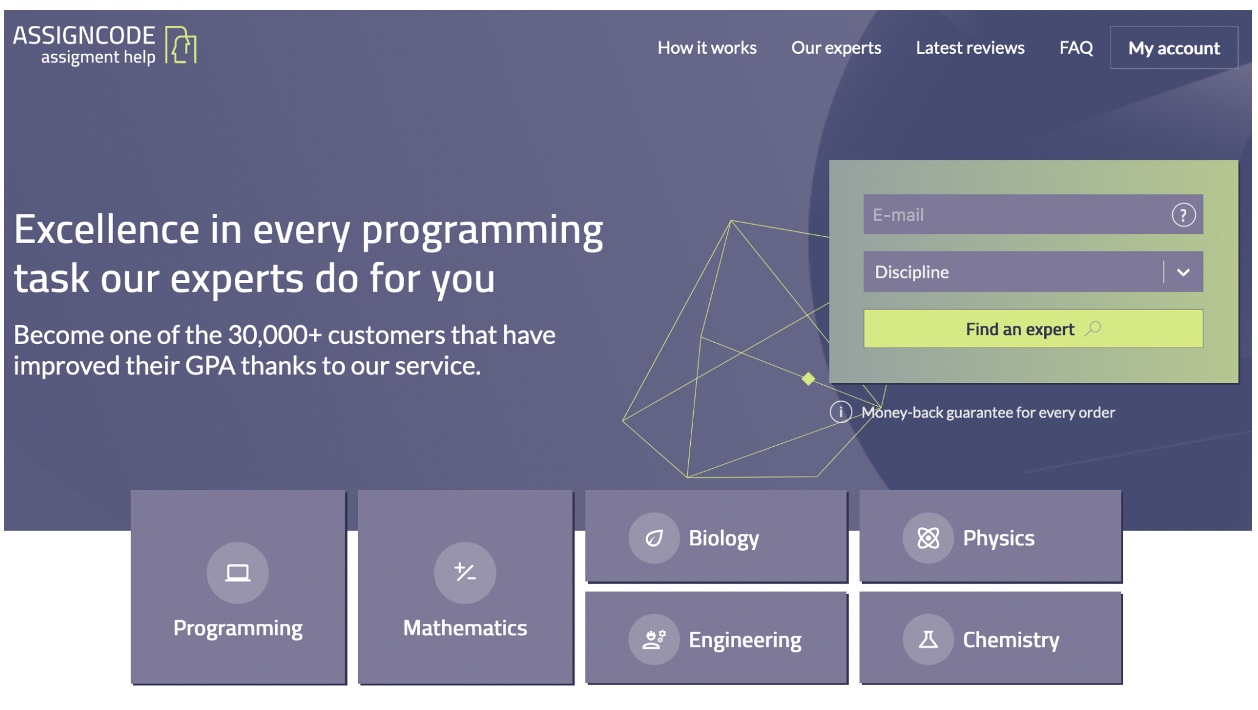
AssignCode.com – A top choice for students who need help with their coding assignments
It’s an ideal website for students looking to get a good grade, build a solid programming foundation, and complete their projects on time. Even though the company has been in operation for a shorter time than other writing companies, it has rapidly grown to be a favorite of college students due to its personalized approach when handling programming papers.
The company boasts of having high-profile programming experts due to the street hiring process. The claim is evident in the quality of the computer programs the experts have managed to send out to the clients, which is the reason for its rapid rise across the ranks of online writing. In addition, the company offers its experts room for growth. For this reason, a team of mentors and quality assurance managers ensures the experts evolve with every rolled assignment. As a result, the group continues to advance in experience as they handle more coding tasks.
AssignCode has been dedicated to offering the best writing experience to its clients. The company has established several guarantees to ensure the students get the best programming assignment help possible. Some of the company’s guarantees include a money-back assurance, customer-oriented service, timeliness, authenticity and confidentiality, and unlimited revision policies. These aspects have made the company stand out in the industry and a good choice for students struggling with coding assignments.
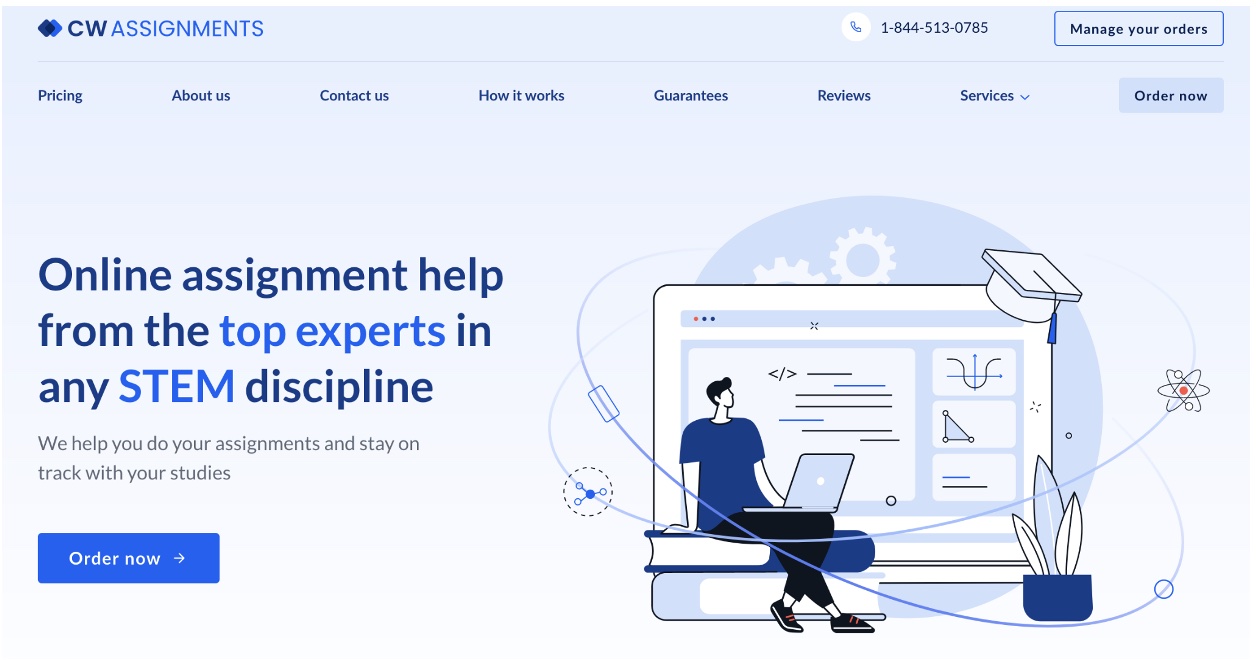
CWAssignments.com – A fantastic resource for college students seeking help with their coding assignments
The website offers a wide range of services, including expert assistance with programming assignments in various languages, such as C++, Java, Python, and more. The website handpicks tutors who are experts in their field and have a proven track record of success in teaching and mentoring students. Therefore, you can be confident that you are getting help from someone who truly knows their stuff when you choose to work with the service.
A unique feature of CWAssignments is its personalized approach to helping students. Instead of simply providing pre-written solutions, the tutors work with you to understand your specific assignment and help you learn how to solve it independently. The custom approach enables you to achieve a better grade on the project and helps you develop the skills you need to succeed in your programming courses. Hence, it’s ideal for college students struggling with complex programming concepts, such as data structures.
In addition, the company has a dedicated and responsive customer service team that is always ready to provide the information you need. They always answer quickly and try to help you as much as possible. The overall experience, based on the numerous online reviews, is that the company has cultivated a culture in which service is customer-oriented. As a result, you will experience an unmatched peace of mind while the experts work on your paper to give you the best results.
CodingPedia.org – A high-speed delivery service for programming and STEM assignments
The website is well-designed and easy to navigate, making it easy to find the information you need. The tutorials on CodingPedia are top-notch, providing clear and concise explanations of various coding theories and example code to help you understand how the concepts are used in solving real-world programming problems. The topics covered on the site are varied and extensive, ranging from basic programming concepts to more advanced topics such as data structures and algorithms.
One thing that sets CodingPedia apart from other coding resources is the quality of the content. The papers are written by experienced developers and are rigorously reviewed to ensure accuracy and clarity. As a result, you can trust that the scripts you receive from the experts are the best and will earn you the best grades. CodingPedia.org papers are priced in different tiers. The flexibility in prices and the consideration of varying delivery timelines set the service apart from all the others. Therefore, the website stands out as an ideal choice when you are looking to have your assignment delivered within the shortest time possible.
There are many reasons you should consider working with CodingPedia.org. First, there are several guarantees to the students. The website’s promises are anchored on four pillars: originality, punctuality, security, and quality. As a programming student and a client, the company guarantees to provide complete programming homework assignments that are free from plagiarism and of the best quality. If the paper still needs to meet your expectations, you’re free to ask for revisions until satisfaction is achieved. In addition, you’re free to choose an expert who will deliver the paper in record time. The level of punctuality within the service has made it one of the options when you’re looking to get your solutions within a short deadline. These features put the company forward as one of the best programming assignment websites on the internet. Therefore, you must not hesitate whenever you need help.
Criteria for Gauging TOP Programming Homework Help Sites for Students in 2023
Several criteria can be used to gauge whether a given programming homework help service is of the best quality. The most significant factors to are:
- Quality of tutors: a top programming homework help site should have a team of highly skilled and experienced tutors who possess a wide range of knowledge and skills in programming languages and disciplines. They should provide students with detailed and accurate answers to their questions.
- User-friendly interface: a good site should have a user-friendly interface that is easy to navigate and understand and allows students to find the resources and help they need quickly.
- Availability: a top site should be available 24/7, so students can get help whenever they need it, regardless of their time zone or schedule.
- Timeliness of delivery: a good site should guarantee that students will receive their assignments on time. Time observance means that students can allocate some time to go through the written code and understand the logic behind its execution process.
- Plagiarism-free guarantee: a high-quality coding assignment help service must understand that plagiarism has severe repercussions for clients. Therefore, the experts must be able to write code from scratch, considering the instructions provided by the student.
- Variety of resources: a good site should provide various resources such as sample assignments, a library, a blog, and a forum where students can connect with other students and share information.
- Flexibility in payment methods and money-back policy: a good site should offer flexible payment options and a money-back policy in case students are not satisfied with the service.
- Positive reviews: a top site should have a good reputation and many positive reviews from past users.
Students who can master these criteria and apply them on the surface of a programming solver online are likely to land services dedicated to helping students get the best grades without focusing more on money.
Frequently Asked Questions
Learn more about finding help with coding homework by checking out the following questions most college students ask.
Where can I get help with programming homework?
The academic writing industry is one of the fast-growing enterprises around the world. For this reason, finding a coding helper doesn’t require rocket science since many organizations offer online services. There are numerous companies providing help with programming assignments for pay. Therefore, all you have to do is do an internet search, and you’ll get some popular programming help sites. However, you don’t have to go through the struggle as this post has shortlisted the top programming homework help websites that can be useful to you.
Can I pay someone to do my programming assignment?
Yes, you can pay someone to do your programming assignment. Various companies have experts with in-depth knowledge of the different programming languages. As a result, they can provide the best coding help whenever you need it. However, you must be careful when going for the services, as some will scam you in the disguise of helping. Therefore, you can opt for the services we have nominated in this article since they are the most reliable and legitimate online programming assignment help services.
Is programming homework help legit?
Programming homework help is legit. Online writing companies employ experts in the industry with the primary aim of providing students with the right knowledge when it comes to handling programming assignments. The service is legal, so you don’t have to worry about anything. Further, you are always free to use the solutions as inspiration in learning.
Summing up
Identifying a reliable and legitimate online programming assignment homework help site takes work. There are a host of factors to consider before settling on any given service, which presents a significant challenge to students just getting started with online coding helpers. Despite the challenge of choosing reliable coding homework help services, customer reviews have stood out as one of the most significant factors that help potential clients choose a service. It has been one of the most critical drivers while evaluating the services alongside other factors such as the price of the papers, quality, and the writers’ proficiency. Therefore, the list presented in this post will change your choices as you seek a coding helper online. We hope the information within this article helps to boost your academic pursuit even as you use online programming experts to handle the most complex assignments.
Advertising disclosure: We may receive compensation for some of the links in our stories. Thank you for supporting LA Weekly and our advertisers.